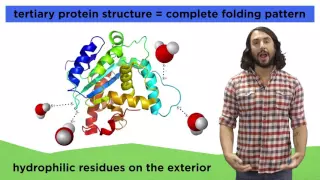
Everyone has heard of proteins. What are they on the molecular level? They're polymers of amino acids, of course. They make up most of your body, so we have to understand their structure very well! Check this out to learn the hierarchy of protein structure so that we can later learn all ab
From playlist Biochemistry

AlphaFold Protein Structure Database
With EMBL-EBI, we're proud to launch the AlphaFold Protein Structure Database, which offers the most complete and accurate picture of the human proteome, doubling humanity’s accumulated knowledge of high-accuracy human protein structures - for free. Try it today: dpmd.ai/alphafolddb
From playlist The story of AlphaFold

Protein Structure - Primary, Secondary, Tertiary, & Quarternary - Biology
This biology video tutorial provides a basic introduction into the four levels of protein structure - primary, secondary, tertiary and quarternary structure. The primary structure of a protein is based on the sequence of amino acids. The secondary structure is based on localized shapes s
From playlist Biochemistry

BERTology Meets Biology: Interpreting Attention in Protein Language Models | AISC
Speaker: Jesse Vig Host: Rouzbeh Afrasiabi Find the recording, slides, and more info at https://ai.science/e/ber-tology-meets-biology-interpreting-attention-in-protein-language-models--f1gSt5qygJhSXf9WG2K1 Motivation / Abstract Transformer architectures have proven to learn useful repres
From playlist ML in Chemistry

The protein folding revolution
Big leaps in our understanding of protein folding can open doors to new protein-based medicines and materials--designed from the ground up. Learn more: http://scim.ag/2a3mGvZ JOIN AAAS: http://scim.ag/2bxrxnH
From playlist News Features

Data structures: Introduction to graphs
See complete series on data structures here: http://www.youtube.com/playlist?list=PL2_aWCzGMAwI3W_JlcBbtYTwiQSsOTa6P In this lesson, we have described Graph data structure as a mathematical model. We have briefly described the concept of Graph and some of its applications. For practice
From playlist Data structures

AMMI 2022 Course "Geometric Deep Learning" - Lecture 12 (Applications & Trends) - Michael Bronstein
Video recording of the course "Geometric Deep Learning" taught in the African Master in Machine Intelligence in July 2022 by Michael Bronstein (Oxford), Joan Bruna (NYU), Taco Cohen (Qualcomm), and Petar Veličković (DeepMind) Lecture 12: What's next? • Beyond traditional Message Passing •
From playlist AMMI Geometric Deep Learning Course - Second Edition (2022)

Lecture 10: What's Next? - Michael Bronstein
Video recording of the First Italian School on Geometric Deep Learning held in Pescara in July 2022. Slides: https://www.sci.unich.it/geodeep2022/slides/Pescara%202022%20-%20conclusions.pdf Blog post: https://towardsdatascience.com/graph-neural-networks-beyond-weisfeiler-lehman-and-vani
From playlist First Italian School on Geometric Deep Learning - Pescara 2022
AMMI Course "Geometric Deep Learning" - Lecture 12 (Applications & Conclusions) - Michael Bronstein
Video recording of the course "Geometric Deep Learning" taught in the African Master in Machine Intelligence in July-August 2021 by Michael Bronstein (Imperial College/Twitter), Joan Bruna (NYU), Taco Cohen (Qualcomm), and Petar Veličković (DeepMind) Lecture 12: What's next? • Beyond Mess
From playlist AMMI Geometric Deep Learning Course - First Edition (2021)
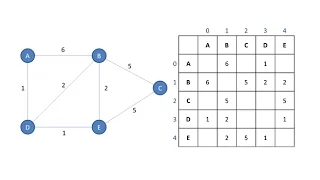
Graph Data Structure 1. Terminology and Representation (algorithms)
This is the first in a series of videos about the graph data structure. It mentions the applications of graphs, defines various terminology associated with graphs, and describes how a graph can be represented programmatically by means of adjacency lists or an adjacency matrix.
From playlist Data Structures
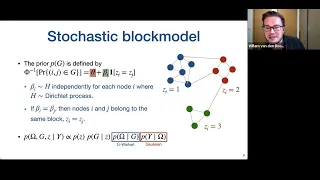

Willem van den Boom - Bayesian Learning of Graph Substructures
Willem van den Boom (National University of Singapore) presents "Bayesian Learning of Graph Substructures, 5 August 2022.
From playlist Statistics Across Campuses

DeepMind's AlphaFold 2 Explained! AI Breakthrough in Protein Folding! What we know (& what we don't)
#deepmind #biology #ai This is Biology's AlexNet moment! DeepMind solves a 50-year old problem in Protein Folding Prediction. AlphaFold 2 improves over DeepMind's 2018 AlphaFold system with a new architecture and massively outperforms all competition. In this Video, we take a look at how
From playlist Papers Explained

Geometric deep learning for functional protein design - Michael Bronstein
Seminar on Theoretical Machine Learning Topic: Geometric deep learning for functional protein design Speaker Michael Bronstein Affiliation: Imperial College London Date: February 20, 2020 For more video please visit http://video.ias.edu
From playlist Mathematics

Proteins in 3D | The Royal Society
We are intrigued by how proteins work. Our genetic code determines the amino acid sequence of proteins, which in turn determines their 3D structure. Subscribe to our channel for exciting science videos and live events, many hosted by Brian Cox, our Professor for Public Engagement: https://
From playlist Latest talks and lectures

David Knowles: "Probabilistic programming for genomics"
Computational Genomics Winter Institute 2018 "Probabilistic programming for genomics" David Knowles, Stanford University Institute for Pure and Applied Mathematics, UCLA February 27, 2018 For more information: http://computationalgenomics.bioinformatics.ucla.edu/programs/2018-cgwi/
From playlist Computational Genomics Winter Institute 2018

Molecular dynamics of the spike protein
Molecular dynamics simulation trajectory of four S proteins embedded in a membrane. The proteins and lipids are shown in surface representation. Glycans are represented by green van der Waals beads. Water and ions are omitted for clarity (simulation time shown: 600ns). More information: ht
From playlist Science Snippets

SDS 429: 2020's Biggest Data Science Breakthroughs — with Jon Krohn
Jon Krohn joins us for a year-end episode about 2020’s biggest data science breakthroughs and for a big podcast announcement for 2021. In this episode you will learn: • Global warming [0:43] • Our big podcast announcement [2:57] • Who is Jon Krohn? [8:15] • Top 3 technological breakthroug
From playlist Super Data Science Podcast

14. Predicting Protein Interactions
MIT 7.91J Foundations of Computational and Systems Biology, Spring 2014 View the complete course: http://ocw.mit.edu/7-91JS14 Instructor: Ernest Fraenkel This lecture is on predicting protein interactions. He discusses structural predictions of protein-protein interactions. He then talks
From playlist MIT 7.91J Foundations of Computational and Systems Biology

Protein structure by X-Ray scattering
This video is about how mathematical formulas apply to the physical model based on biological purposes. Proteins are the constructor of life. They do almost all functions in us. And this is extremely important to know their structure. Example protein https://en.wikipedia.org/wiki/P53#/m
From playlist Summer of Math Exposition Youtube Videos
